DNA Binding Motif
| Accessions: | 1zg1_AB (3D-footprint 20231221) |
| Names: | Nitrate/nitrite response regulator protein narL |
| Organisms: | Escherichia coli, strain K12 |
| Libraries: | 3D-footprint 20231221 1 1 Contreras-Moreira B. 3D-footprint: a database for the structural analysis of protein-DNA complexes. Nucleic acids research 38:D91-7 (2010). [Pubmed] |
| Description: | NarL complexed to nirB promoter non-palindromic tail-to-tail DNA site |
| Length: | 16 |
| Consensus: | TACTnnTnnaTnGGTA |
| Weblogo: | 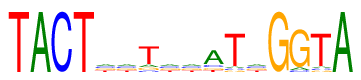 |
| PSSM: | P0 A C G T 01 0 0 0 96 T 02 96 0 0 0 A 03 0 96 0 0 C 04 0 0 0 96 T 05 24 24 24 24 n 06 24 24 24 24 n 07 11 7 11 67 T 08 24 24 24 24 n 09 24 24 24 24 n 10 62 12 11 11 a 11 5 5 5 81 T 12 24 24 24 24 n 13 0 0 96 0 G 14 7 5 84 0 G 15 5 5 5 81 T 16 96 0 0 0 A |
| Binding TFs: | 1zg1_A / 1zg1_B (Bacterial regulatory proteins, luxR family, Sigma-70, region 4, Sigma-70, region 4, Winged helix-turn-helix DNA-binding) |
| Binding Sites: | 1zg1_C 1zg1_D |
| Publications: | Maris AE., Kaczor-Grzeskowiak M., Ma Z., Kopka ML., Gunsalus RP., Dickerson RE. Primary and secondary modes of DNA recognition by the NarL two-component response regulator. Biochemistry. 44(44):14538-52 (2005). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.