DNA Binding Motif
| Accessions: | MA0265.1 (JASPAR 2024) |
| Names: | ABF1 |
| Organisms: | Saccharomyces cerevisiae |
| Libraries: | JASPAR 2024 1 1 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | PBM, CSA and/or DIP-chip |
| Length: | 16 |
| Consensus: | yCGTakrwartGAyaw |
| Weblogo: | 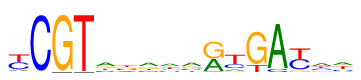 |
| PSSM: | P0 A C G T 01 11 41 4 44 y 02 0 99 0 1 C 03 0 0 99 0 G 04 0 0 0 99 T 05 41 17 22 20 a 06 22 24 26 28 k 07 39 16 28 17 r 08 34 18 20 28 w 09 40 21 24 15 a 10 41 0 57 0 r 11 12 25 9 55 t 12 0 0 81 19 G 13 92 8 0 0 A 14 0 45 9 46 y 15 37 22 18 23 a 16 34 18 14 34 w |
| Type: | Heterodimer |
| Binding TFs: | P14164 (BAF1 / ABF1 chromatin reorganising factor) P14164 |
| Publications: | Badis G, Chan E.T, van Bakel H, Pena-Castillo L, Tillo D, Tsui K, Carlson C.D, Gossett A.J, Hasinoff M.J, Warren C.L, Gebbia M, Talukder S, Yang A, Mnaimneh S, Terterov D, Coburn D, Li Yeo A, Yeo Z.X, Clarke N.D, Lieb J.D, Ansari A.Z, Nislow C, Hughes T.R. A library of yeast transcription factor motifs reveals a widespread function for Rsc3 in targeting nucleosome exclusion at promoters. Molecular cell 32:878-87 (2008). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.