DNA Binding Motif
| Accessions: | MA0258.2 (JASPAR 2024) |
| Names: | ESR2 |
| Organisms: | Homo sapiens |
| Libraries: | JASPAR 2024 1 1 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | ChIP-seq |
| Length: | 15 |
| Consensus: | rGGTCAsmstGaCCy |
| Weblogo: | 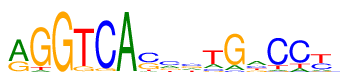 |
| PSSM: | P0 A C G T 01 5410 96 2563 174 r 02 429 0 7170 644 G 03 59 0 8143 41 G 04 74 147 989 7033 T 05 0 7621 533 89 C 06 8098 1 57 87 A 07 502 4202 2738 801 s 08 2092 3235 1688 1228 m 09 1808 2510 2173 1752 s 10 1602 975 533 5133 t 11 1121 398 6658 66 G 12 3278 2056 1391 1518 a 13 434 6469 363 977 C 14 1111 5854 0 1278 C 15 834 2708 606 4095 y |
| Type: | Heterodimer |
| Binding TFs: | Q92731 (Ligand-binding domain of nuclear hormone receptor, Zinc finger, C4 type (two domains), Estrogen receptor beta) Q92731 |
| Binding Sites: | MA0258.2.1 MA0258.2.10 MA0258.2.11 MA0258.2.12 MA0258.2.13 MA0258.2.14 MA0258.2.15 MA0258.2.16 MA0258.2.17 MA0258.2.18 MA0258.2.19 MA0258.2.2 MA0258.2.20 MA0258.2.3 MA0258.2.4 MA0258.2.5 MA0258.2.6 MA0258.2.7 MA0258.2.8 MA0258.2.9 |
| Publications: | Liu Y, Gao H, Marstrand T.T, Ström A, Valen E, Sandelin A, Gustafsson J.A, Dahlman-Wright K. The genome landscape of ERalpha- and ERbeta-binding DNA regions. Proceedings of the National Academy of Sciences of the United States of America 105:2604-9 (2008). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.