DNA Binding Motif
| Accessions: | PAX7_full (HumanTF 1.0), MA0680.1 (JASPAR 2024) |
| Names: | PAX7 |
| Organisms: | Homo sapiens |
| Libraries: | HumanTF 1.0 1, JASPAR 2024 2 1 Jolma A, Yan J, Whitington T, Toivonen J, Nitta KR, Rastas P, Morgunova E, Enge M, Taipale M, Wei G, Palin K, Vaquerizas JM, Vincentelli R, Luscombe NM, Hughes TR, Lemaire P, Ukkonen E, Kivioja T, Taipale J. DNA-Binding Specificities of Human Transcription Factors. Cell. 2013 Jan 17;152(1-2):327-39. [Pubmed] 2 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | Site type: dimeric; SELEX cycle: 3, HT-SELEX |
| Length: | 10 |
| Consensus: | TAATCRATTA |
| Weblogo: | 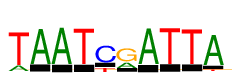 |
| PSSM: | P0 A C G T 01 679 68 98 3708 T 02 3708 18 16 6 A 03 3708 3 0 0 A 04 6 21 3 3708 T 05 10 2934 24 774 C 06 1135 27 2573 0 R 07 3708 0 10 10 A 08 12 0 0 3708 T 09 21 25 9 3708 T 10 3708 72 47 249 A |
| Type: | Heterodimer |
| Binding TFs: | PAX7_full (Homeobox domain, 'Paired box' domain, Homeobox KN domain, Paired box protein 7 ) P23759 (Homeobox domain, 'Paired box' domain, Homeobox KN domain, Paired box protein 7 ) P23759 |
| Publications: | Jolma A, Yan J, Whitington T, Toivonen J, Nitta KR, Rastas P, Morgunova E, Enge M, Taipale M, Wei G, Palin K, Vaquerizas JM, Vincentelli R, Luscombe NM, Hughes TR, Lemaire P, Ukkonen E, Kivioja T, Taipale J. DNA-Binding Specificities of Human Transcription Factors. Cell. 2013 Jan 17;152(1-2):327-39. [Pubmed] Smith S. B., Ee H. C., Conners J. R., German M. S. Paired-homeodomain transcription factor PAX4 acts as a transcriptional repressor in early pancreatic development. Mol. Cell. Biol. 19:8272-8280 (1999). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.