| structure name | CRYSTAL STRUCTURE OF THE BACTERIOPHAGE MU TRANSPOSOSOME (TRANSPOSASE) |
| reference | Montano et al. Nature 491 413 2012 |
| source | ENTEROBACTERIA PHAGE MU |
| experiment | X-ray (resolution=3.71, R-factor=0.395) |
| structural superfamily | Ribonuclease H-like;Homeodomain-like;mu transposase, C-terminal domain; |
| sequence family | Bacteriophage Mu transposase; Mu DNA binding, I gamma subdomain; Mu transposase, Cterminal; |
| multimeric complexes | 4fcy_AB |
| links to other resources | NAKB  PDIdb DNAproDB PDIdb DNAproDB |
| protein sequence | |
| interface signature | STRDYRSRAF |
Estimated binding specificities ?
readout + contact + contact | 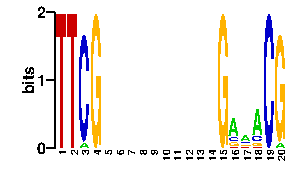 |
A | 5 0 16 16 16 0 24 24 24 24 24 24 24 24 24 24 0 5 96 96 C | 90 0 5 15 15 91 24 24 24 24 24 24 24 24 24 24 96 0 0 0 G | 1 96 10 15 5 0 24 24 24 24 24 24 24 24 24 24 0 91 0 0 T | 0 0 65 50 60 5 24 24 24 24 24 24 24 24 24 24 0 0 0 0scan! |
Dendrogram of similar interfaces ?
matrix format------SCTGRTDCYTRGSGRAAA--FT-------------------- L +---------------------4fcy_B +---2 ------------------------------WA--TA--ECRT--RC-- L +-3 +---------------------7jn3_E ! ! ------------------------------------QGECSG--HCQT +-4 +-------------------------6rwm_A ! ! ----------------------------------------ECSC--KG ! ! +--6vdk_A +-5 +-----------------------1 --------------------------------QT----ECRG--QG-- L ! ! +--7u32_A --6 ! HGSGRG--HG--RT----STKGRGTTLGNT------------------ L ! +----------------------------1u78_A ! --SGRGPT--SGRG---------------------------------- L +----------------------------7bhy_A |
home
updated Tue Dec 19 08:46:13 2023
