| structure name | RECOGNITION OF AT-RICH DNA BINDING SITES BY THE MOGR REPRESSOR (Motility gene repressor mogR) |
| reference | Higgins et al. Structure 17 769 2009 |
| source | Listeria monocytogenes |
| experiment | X-ray (resolution=1.75, R-factor=0.217) |
| multimeric complexes | 3fdq_AB |
| links to other resources | NAKB  PDIdb DNAproDB PDIdb DNAproDB |
| protein sequence | |
| interface signature | VSQNYPR |
Estimated binding specificities ?
| readout + contact | 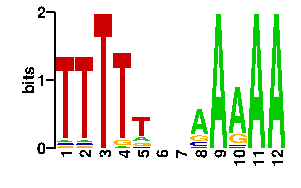 |
A | 0 0 0 0 13 24 24 73 68 89 85 C | 0 0 24 0 18 24 24 0 27 0 1 G | 0 0 4 0 24 24 24 6 0 4 0 T | 96 96 68 96 41 24 24 17 1 3 10scan! |
Related DNA sequences reported in the literature ?
| site | source | matches (E-value) |
|---|---|---|
| term: MOGR | ||
| ATTTTTTAAAAAAAT | PubMed | 3fdq_A(1.28e-11) |
Dendrogram of similar interfaces ?
matrix format------------------VT------SAQTNAYT----PTRT L +-----------------3fdq_A +-----1 ------------RC--RT------------NCSTNARG---- L +-2 +-----------------1mnm_C ! ! NT--NAITRG-------------------------------- +-3 +-----------------------5zjt_B ! ! --------------RT--------STVCKGDCRG-------- L +-4 +-------------------------1w0t_A ! ! IAQCNAYA--RT------------------------------ L +-5 +--------------------------1nk2_P ! ! ------------------------RCET--SAQAKT------ L +-6 +---------------------------6qec_A ! ! VAQTNA------------------------------------ L +-7 +---------------------------5hod_A ! ! --------------------HGSGRGHGRT------------ L --8 +----------------------------1tc3_C ! NT--NAIARG-------------------------------- L +----------------------------1b72_B |
home
updated Tue Dec 19 07:19:07 2023
