| structure name | CRYSTAL STRUCTURE OF THE METHYLATED N-ADA/DNA COMPLEX (Ada polyprotein) |
| reference | Norman et al. Mol.Cell 20 117 2005 |
| source | Escherichia coli |
| experiment | X-ray (resolution=2.10, R-factor=0.235) |
| structural superfamily | Ada DNA repair protein, N-terminal domain N-Ada 10; |
| sequence family | Metal binding domain of Ada; Bacterial regulatory helixturnhelix; |
| redundant complexes | 1zgw |
| reference complex | 1zgw_A |
| links to other resources | NAKB  PDIdb DNAproDB PDIdb DNAproDB |
| protein sequence | |
| interface signature | TTRRF |
Estimated binding specificities ?
| readout + contact | 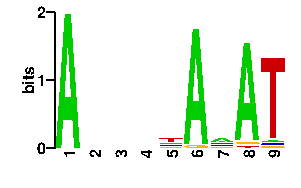 |
A | 90 0 18 6 54 24 24 24 0 C | 6 6 24 0 12 24 24 24 0 G | 0 0 12 6 12 24 24 24 0 T | 0 90 42 84 18 24 24 24 96scan! |
Dendrogram of similar interfaces ?
matrix formatTTTTRTRT----FT------ L +----------------1u8b_A +------1 --------DTRTRGTTKT-- L +-2 +----------------2o8k_A ! ! ------------------YG +-3 +-----------------------1sfu_A ! ! ----------------RGYG --4 +-------------------------1qbj_C ! ------------------YG +--------------------------3irr_C |
home
updated Mon Dec 18 06:11:39 2023
